Principali standard e linee guida e come le attività del CRS4 contribuiscono al loro miglioramento.
In questo focus Alessandro Sulis (CRS4, gruppo Healthcare Flows), parla di interoperabilità tra sistemi clinici, illustrandone i principali standard e linee guida e descrivendo in che modo le attività del CRS4 contribuiscono al loro miglioramento.
In un ospedale moderno è essenziale che i diversi domini clinici (laboratorio, radiologia, ecc.) siano in grado di scambiarsi informazioni. Tecnicamente, questo implica che differenti domini clinici siano in grado di scambiarsi dei messaggi mutuamente comprensibili, i quali trasportano informazioni relative a pazienti e processi clinici. Questo requisito, apparentemente semplice, è difficile da raggiungere nella pratica. Infatti, i sistemi clinici tendono a essere progettati come “isole informatiche” separate, le quali non comunicano facilmente col resto del mondo.
Per ovviare a questo, negli ultimi vent’anni sono stati sviluppati diversi standard e linee guida, che attualmente sono sempre più supportati dai vendors. Il CRS4 contribuisce attivamente allo sviluppo di questi standard e linee guida – in particolare quelli relativi ai domini clinici del Laboratorio e della Anatomia Patologica – ed alla loro implementazione in contesti clinici reali.

Alessandro Sulis, Laurea in Ingegneria Elettronica all'Università di Cagliari e Master di II livello in Tecnologie dell'Informazione al CEFRIEL di Milano, lavora al CRS4 dal 2007 e attualmente si occupa di interoperabilità̀ tra sistemi clinici secondo i principali standard e linee guida, e collabora attivamente con gli organismi internazionali di riferimento (IHE e HL7 International); in particolare, è membro del Technical Committee del dominio PaLM (Pathology and Laboratory Medicine) di IHE.
— E. Falqui
Technical Corner
Health IT interoperability has an important role in health service quality and effectiveness at different levels, from the intra-institutional communication between department-based clinical systems (such as Laboratory Information Systems, Radiology Information Systems, etc.) to the inter-institutional interaction and coordination of regional or national components of a healthcare system. Also, in clinical and biomedical research, data integration is essential as a basis for guiding differential outcome analysis. Interoperability implies an exchange and mutual interpretation of information by two or more systems. In fact, the first definition of interoperability in the IEEE Standard Computer Dictionary stated, “the ability of two or more systems or components to exchange information and to use the information that has been exchanged”1. Notice that the concept is not limited to information exchange, since this action is necessary but not sufficient to achieve interoperability. Focusing on health informatics, HL7 International2 gave a more detailed characterization3 identifying three main types of health IT interoperability:
- technical interoperability, focusing on physical communication of data;
- semantic interoperability, relating to the meaning of data exchanged;
- process interoperability, addressing systems integration into a work setting.
This classification goes beyond simple labels, creating a hierarchical model which can guide interoperability implementation in different contexts: the activity of Healthcare Flows Research Program spans all the three types above and the following paragraphs show some of the results obtained in this field.
Technical Interoperability
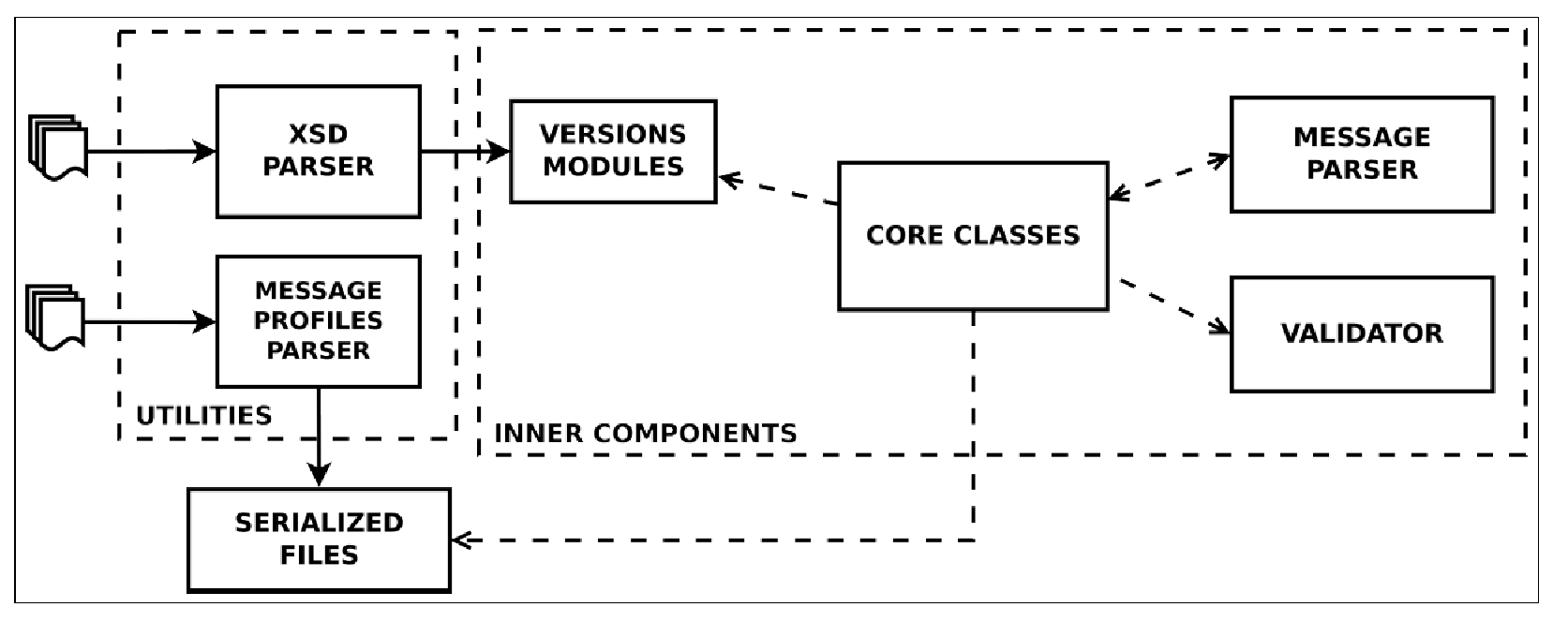
From the technical interoperability perspective, the group develops and maintains HL7apy, an innovative Python package to manage HL7 v2.x messages for fast prototyping of HL7-compliant software. As shown in Figure 1, the main library components are:
- inner components: core classes and an API to create and manage HL7 messages; Message Parser, to parse ER7-encoded messages; and Validator, to validate messages;
- two utility scripts: an XSD Parser, to generate Python modules for every HL7 v2 minor version; and a Message Profiles Parser, to serialize files for message profiles.
The main classes defined in HL7apy to represent HL7 elements are:
- Messages
- Group
- Segment
- Field
- Component
- SubComponent
- Base datatype classes
The core classes allow the developer to create HL7-compliant messages, to navigate their structure and to manipulate HL7 elements in a compact form, as shown in the code examples in Figure 2, Figure 3 and Figure 4.
One of the most important features provided by HL7apy is the validation of messages, since it ensures their compliance with the standard for the specific message type and HL7 version. The library allows two levels of validation: STRICT and TOLERANT.
A detailed description of HL7apy can be found in4; the project is still under development and the code is released under an open-source license5.
Semantic and Process interoperability
The Healthcare Flows group also actively works on semantic and process interoperability, collaborating intensely with IHE6– the reference institution for information sharing and healthcare systems interoperability. In 2010 the group joined the IHE Technical Committee for the Laboratory Domain, now called “Pathology and Laboratory Medicine (PaLM)” Domain. Our first contribution in this context was the proposal of an extension to the IHE Laboratory guidelines to enhance traceability in the pre-analytical phase, going from the prescription of exams to sample preparation for analysis7. For each domain, IHE models clinical processes by defining the main workflow steps in detail, the main use cases, the actors involved, the transactions and, for each transaction, the messages to be used and the specific meaning of each message field. Concerning the laboratory pre-analytical process, IHE defines two main steps, identification and sample collection, mapping them to IHE PDQ (Patient Demographics Query) and LBL (Laboratory Specimen Barcode Labeling) profiles. The actors involved are:
- Patient Demographics Consumer (PDC): to query a set of patient information;
- Patient Demographic Supplier (PDS): provides patient-specific information;
- Label information Provider (LIP): the actor that provides information about labels;
- Label Broker (LB): the actor responsible for the labeling of the containers according to the information provided by the LIP.
The transactions for the two profiles are:
- ITI-21 and ITI-22 for PDQ;
- LAB-61 and LAB-62 for LBL
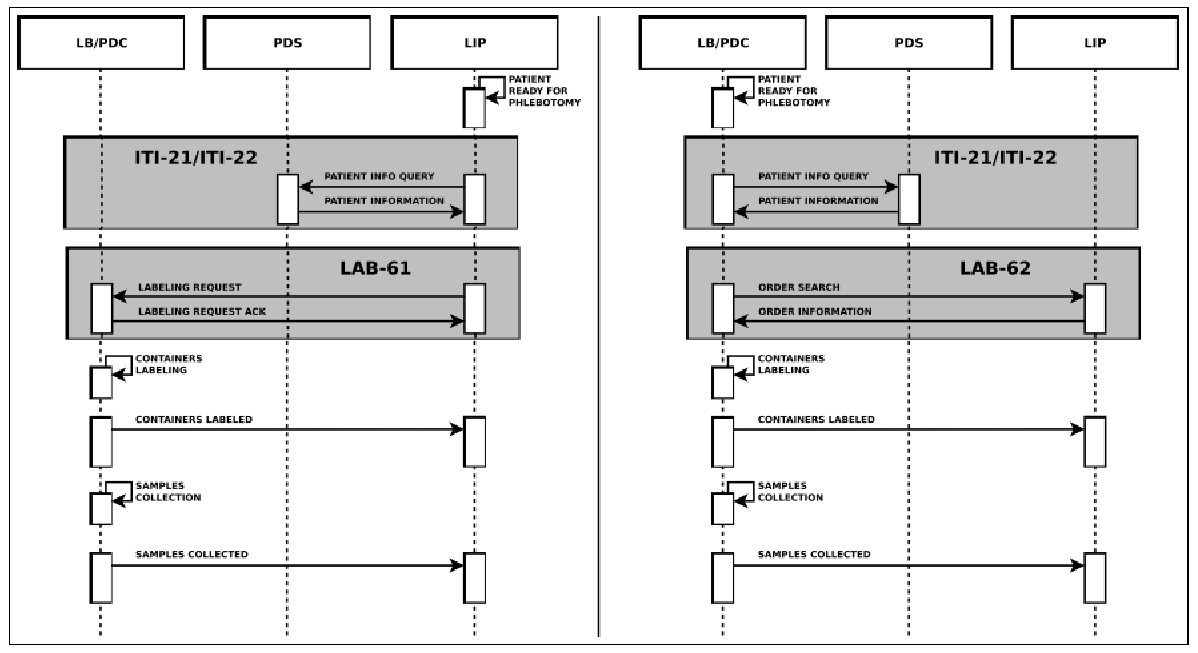
All of them are based on HL7 messages exchange. Concerning LBL, in a phlebotomy process supported by automation, the robotized labeling of specimen containers considers two different scenarios: Request Mode and Query Mode. These are both showed in Figure 5.
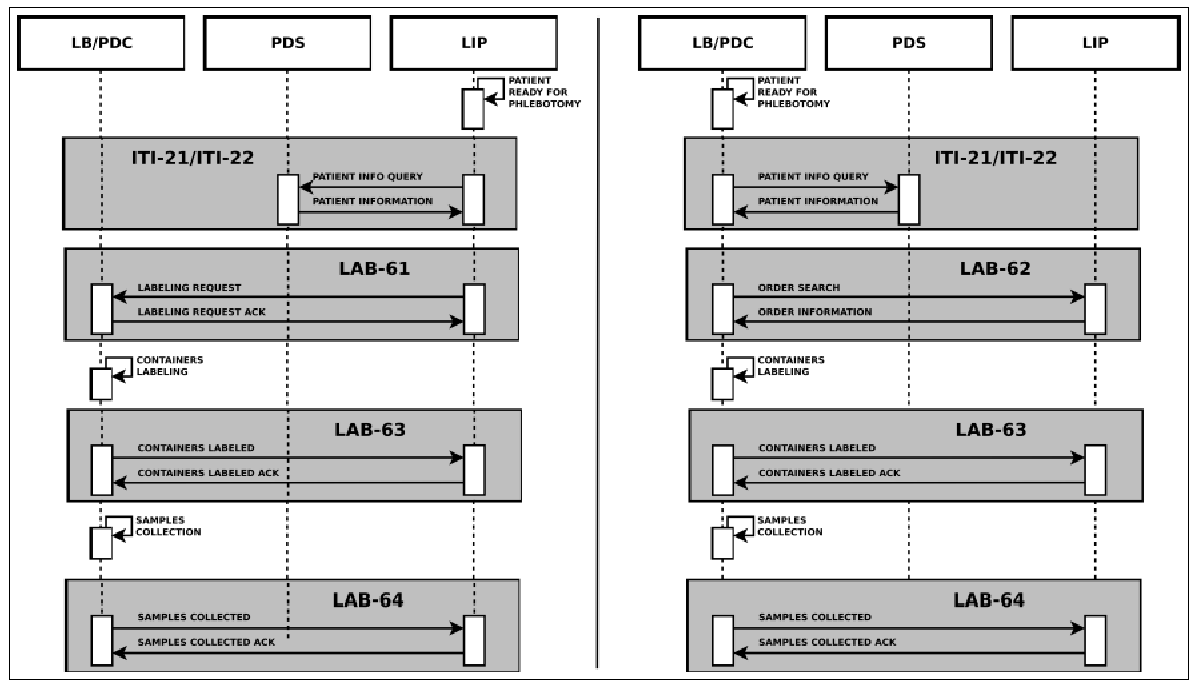
The main information needed for the labeling process is carried by the HL7 messages exchanged between the actors. In detail they provide: patient data, drawn specimens with their unique id, tests to be performed on every specimen and type of container to use. This information is very useful for traceability purposes, but no details are provided by the LBL profile regarding the preparation of the sample container and the sample collection. For these reasons, we proposed an extension of this profile, adding two new transactions, LAB-63 (Labeled Containers Production Confirmation) and LAB-64 (Specimen Collection Confirmation), depicted in Figure 6.
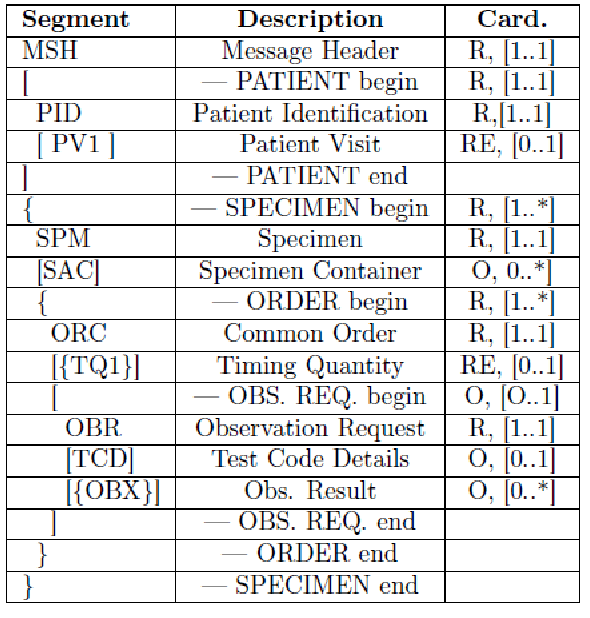
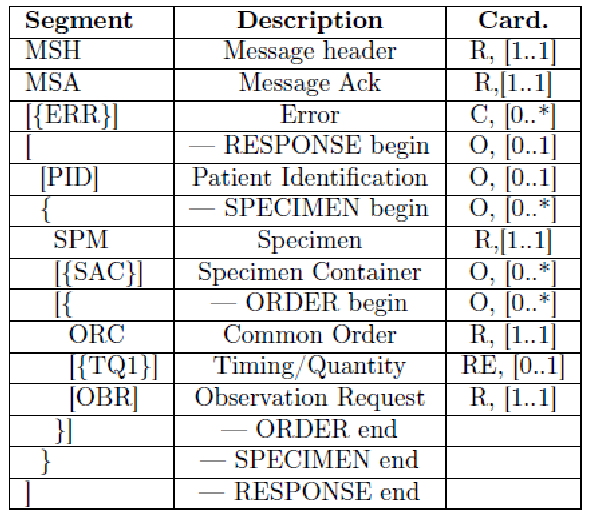
The HL7 messages chosen to define these transactions are OML^O33 (Laboratory Order Message) and ORL^O34 for acknowledgment, since they are specimen-centric and their structures contain all the information needed for traceability purposes.
We submitted the profile extension in a Supplement to the Committee for public discussion in July 2011. LAB-63 was reviewed and immediately accepted and – after the trial implementation period – was included in IHE PaLM Domain Technical Framework8. LAB-64 was considered above LBL scope, as it also involves other IHE profiles; for these reasons, the proposed profile became the starting point for a completely new profile, called Specimen Event Tracking (SET), which we have been developing since 2016 in strict collaboration with PALM Technical Committee.
http://www.en13606.org/the-ceniso-en13606-standard/semantic-interoperability ↩
HL7 EHR Interoperability Work Group, “Coming to terms”, available at https://www.hln.com/assets/pdf/Coming-to-Terms-February-2007.pdf ↩
Meloni V. et al, “HL7apy: a Python library to parse, create and handle HL7 v2.x messages” ”, EJBI - European Journal for Biomedical Informatics, Volume: 12, 2016, available at http://www.ejbi.org/scholarly-articles/hl7apy-a-python-library-to-parse-create-and-handle-hl7v2x-messages.pdf ↩
Sulis A. et al, “Traceability Based Description of Clinical Processes: Extension of IHE Guidelines for Phlebotomy Workflows”, EJBI - European Journal for Biomedical Informatics, Volume: 12, 2016, available at http://ihic2016.eu/Doc/02_2016_06_13_HL7apy@IHIC_Sulis.pdf ↩